Platform news
This page lists significant improvements to the platform and other important updates since June 2021, with the most recent at the top. We suggest you check it regularly.
Last updated: 01 March 2024
New ordering of workspaces on OpenSAFELY Jobs homepage
Updated: 01 March 2024
What has changed?
Once logged into OpenSAFELY Jobs, the 5 workspaces shown to you on your home page are now the five most recently active (by creation/modification date or most recently run job) that you have access to. This is a change from previously showing you the five most recently created.
What does this mean for me?
For most users with a small number of workspaces, this change in ordering will likely have little effect. For those with access to many workspaces, this will hopefully make it easier to get access to the most relevant workspaces as quickly as possible.
- Pull request: opensafely-core/job-server#4165
Comparing study code on OpenSAFELY Jobs
Updated: 28 February 2024
OpenSAFELY Jobs has always shown the Git commit hash for a given job request or job, with a link to view the workspace’s code on GitHub as of that revision. We now also provide GitHub code comparison links (example) so you can see exactly what code has changed between this revision and previous versions.
Job Requests
The code comparison links can be found on a job request’s page in the Code comparison section beneath the job information. If this is the first job request for this workspace on this backend, no code comparison is possible and so no link is available. If there are previous job requests, a link to compare the workspace’s code for this job request to the previous, and previous successful runs (if present) will be present.
Jobs
The code comparison links can be found on a job’s page at the bottom of the Job information section. If this is the first run of this action in this workspace on this backend, no code comparison is possible. If it has been run before, links to compare the workspace’s code for this job to the previous, and previous successful runs (if present) will be present.
N.B. If there lots of changes between versions of a workspace’s
codebase, the code comparisons for a job may also include changes
that are not directly related to the job in question. Identification
of the job by its action name in the project.yaml file and
examination of changes to related files listed in its configuration
should reveal what the relevant changes are.
- Pull request: opensafely-core/job-server#4130
Markdown formatting of Project Status Description
Updated: 26 February 2024
Users can now use Markdown formatting to update the Status Description field in the Project page.
Screenshots


Example URLs
ehrQL v1 released
Updated: 11 December 2023
We have released v1 of ehrQL.
It contains a small number of breaking changes from v0.
For existing users of ehrQL, please refer to the release notes for guidance about updating from v0.
Thank you to all users who have tried out ehrQL and given us feedback!
ehrQL’s measures framework has disclosure control enabled by default
Updated: 27 November 2023
ehrQL’s measures framework is used to calculate quotients (i.e. a numerator divided by a denominator) and to see how these vary over time and when broken down by different groupings.
Previously, numerators and denominators were not subject to disclosure control: a user had to apply disclosure control by writing their own downstream action. Now, ehrQL’s measures framework has disclosure control enabled by default. First, values less than or equal to seven are replaced with zero (suppressed); then, values are rounded to the nearest five.
When the built-in disclosure control method is unhelpful,
such as when an alternative method is required or when unsuppressed/unrounded values are needed by a downstream action,
the configure_disclosure_control method
allows a user to disable disclosure control.
For example:
measures = create_measures()
measures.configure_disclosure_control(enabled=False)
measures.define_measure(...)
For more information about disclosure control in OpenSAFELY, please see the “Updated disclosure control guidance” page.
As always, if you need help with ehrQL, then please ask for help on the #ehrql-support Slack channel. (If you’re unsure how to join, then please ask your co-pilot.)
Warn about invalid moderately_sensitive outputs
Updated: 14 November 2023
The opensafely tool has been
updated to warn you if you have invalid outputs marked as
moderately_sensitive in your project.yaml. This is a follow on from the
previous change about stricter output file
paths and means that you’ll
get more accurate feedback when running the code locally.
Specifically, it will check that moderately_sensitive outputs meet the
appropriate output file
restrictions.
These currently are:
The file must be of the correct type. You will not be able to run jobs locally at all or on the server if the file is not a valid type, i.e. it must have a valid file extension.
If it is a
.csvfile, it must not have apatient_idcolumn. Your code will still run, but the log file and the on-screen summary text will show a warning. If you run it on your own computer using the opensafely command line tool, you will still get an output. If you run it in the live system via jobs.opensafely.org, then it will still run, but the file will not be available in level 4If it is too large, it will be handled in the same way as above. This is unlikey to occur when running locally against dummy data, but may happen when run via jobs.opensafely.org.
Fixing these is likely a case of marking the file as highly_sensitive
instead. If you do need it to be moderately_sensitive, then you may need to
process the data a bit more, e.g. remove the patient_id column or reduce the
size.
As a reminder, the policy for moderately_sensitive senstitive outputs is that
they must be aggregate data, not patient
level.
These checks are designed to catch accidental misclassification of outputs with patient level data
as moderately_sensitive.
Any questions or problems, please let us know.
Stricter output paths in project.yaml
Updated: 07 November 2023
We’ve made some changes to slightly restrict output file paths.
Specifically, all paths must now end with file extension, e.g. .csv.
Previously, it was possibly to use a trailing * character to match all files
(e.g. outputs/data.*). You can still use * to match groups of similar
files, but you must end the pattern with a file extension (e.g.
outputs/data*.csv).
Nearly all project.yaml files do already have explicit file extensions in all their output paths, and this will not require any changes for most users. There are a few users who may need to add an explicit file extensions in order for jobs to run, both locally and on backends. The opensafely tool that you run locally has been updated, and it will warn you that you need to do this.
There are multiple motivations for this change, but the main one is that we
want to be able to inspect the file types when loading a project.yaml. This
allows us to warn users if they are trying to mark an unsupported file
type
as moderately_sensitive.
It also stops accidentally capturing output files that were not intended to be outputs (which incurrs a performance and disk space cost).
Any questions or problems, please let us know.
Updated packages in R image
Updated: 22 September 2023
In order to add a new package to the R Docker image we had to upgrade some of the existing package versions. Hopefully these upgrades will be welcome but there’s a small possibility they might cause issues with existing code so please be on the look out for this.
The most significant upgrade is of the readr package from 1.3.1 to 2.1.4.
You can find the changelog here:
https://readr.tidyverse.org/news/index.html
You can see the full list of upgraded packages on Github here: https://github.com/opensafely-core/r-docker/commit/30fc019
Automatically run failed dependencies when submitting jobs
Updated: 19 September 2023
Previously, when submitting jobs to run the platform would reject requests if any of the requested action’s dependencies had previously failed.
This required users to explicitly request those failed dependencies to be
re-run. This was cumbersome, and was not the same behaviour as when running
actions locally on user’s machines with the opensafely run command.
We have now changed this behaviour, so that if you submit an action to run, any dependent actions that have previously failed will also be scheduled to run. This should make submitting jobs simpler in many cases.
For context, the original design for this behaviour was to encourage users to ensure their code runs succesfully locally before submitting to run for real. This avoids overloading the system with jobs that may fail anyway due to code errors. This is still a very important best practice what we strongly encourage users to do before submitting jobs.
However, this resulted in an awkward workflow for submitting batches of jobs, hence this change.
Updated disclosure control guidance
Updated: 18 September 2023
We have recently made some updates to our disclosure control guidance. This is important for both researchers requesting release of outputs and those who review them. We have made the following changes, which are summarised in more detail below:
- Requiring rounding of counts in addition to suppression of low counts
- Increasing the small number suppression threshold to 7
- Encouraging release of underlying data earlier in the analysis pipeline
- A new checklist for requesting a release
- Updated recommendations for release of log files
- More detail on allowed file types
User, project, and organisation Event Logs added
Updated: 05 September 2023
OpenSAFELY Jobs has been updated with new Event Log pages for users, projects, and organisations.
These pages provide specific information about current Job Requests, and allow users to see a complete audit log of all previous Job Requests.
Examples of new Event Log pages include:
- User: Louis Fisher
- Organisation: The London School of Hygiene & Tropical Medicine
- Project: Investigating the effectiveness of the COVID-19 vaccination programme in the UK
The global OpenSAFELY Job Server Event Log page has also been updated to match the other Event Log pages, and contains a complete log of all Job Requests ever run by OpenSAFELY across all Backends.
As a user of the Job Server, you can access your own Event Log page from your dashboard.
New opensafely clean command
Updated: 12 April 2023
We have added a command to the opensafely command line tool: opensafely clean
This command will safely remove any leftover OpenSAFELY docker artifacts from your system. Specifically, it removes old images, job containers, and file volumes that may have been created when running jobs locally on a user’s computer.
Users who have Docker runnng on WSL may find this particularly useful, as it looks like removing these leftover artifacts can help reduce memory usage.
The documentation has more information on managing your local docker resources.
Improved access to OpenSAFELY Jobs from secure environments
Updated: 06 April 2023
OpenSAFELY Jobs is used by researchers from within the secure environments (such as TPP level 4) to support the review and output checking of files.
We’ve added an easier way for users to log in to OpenSAFELY Jobs from the TPP secure environment, by implementing auto-generated single use tokens. Previously, users were required to manually type their Github password and 2FA token, which was too easy to get wrong.
To use this new method, users first generate a single use token by going to their settings page and clicking on the “Generate Single Use Token” button. Then, they log in as normal to TPP level 4 over the VPN and open a browser window pointing at the OpenSAFELY Jobs log in page. They are now able to authenticate to OpenSAFELY Jobs using just their username or email and the single use token. Much quicker while still being highly secure!
Updates to the OpenSAFELY Jobs home page
Updated: 27 March 2023
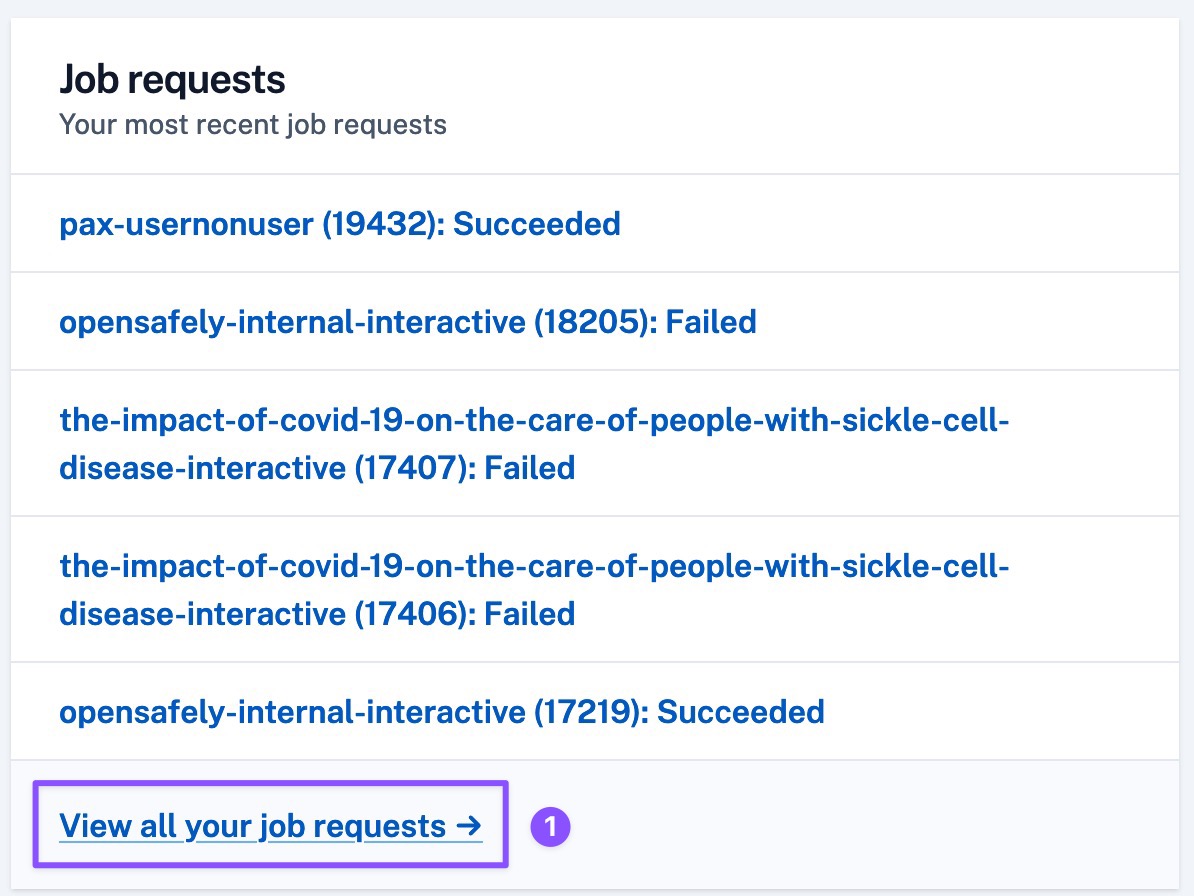
OpenSAFELY Jobs home page has been updated to create a personalised dashboard for users.
Once logged in, users will now be able to view:
- The most recent job requests they have created
- Workspaces they have access to
- Most recently updated projects
- In progress and completed application forms
Alongside the new home page, users are now able to access a personalised list of:
For visitors to the website who don’t have an account, a new home page is visible containing information on why they might want to use OpenSAFELY for their next research project.
Introducing the opensafely exec command for interactive sessions.
Updated: 16 February 2023
New opensafely command: opensafely exec
There is new opensafely exec command in recent versions of the opensafely
tool. It is designed to aid in development of analysis code using the published
OpenSAFELY docker images.
The need to add your your work-in-progress code to project.yaml in order to test it is awkward. This leads some users to use their own locally installed R/python/stata tooling to develop their code. However, this means they may end up inadventantly using a library (or version of library) which is not available in the relevant OpenSAFELY image, and then the code fails when running with project.yaml.
We previously added the opensafely jupyter
command
to help improve this workflow when working with Jupyter notebooks, but with
opensafely exec, we’ve extended this support to other tools.
What does it do?
Running opensafely exec IMAGE COMMAND does the following:
- runs the relevant docker IMAGE (r, python, stata-mp, cohortextractor)
- shares the files in your current directory
- executes COMMAND (or the default command for the image if you don’t supply one)
This might sound fairly simple, but it opens up some convenient new workflows.
Examples
To run an interactive R session in your current directory, just do:
opensafely exec r R
This will start the opensafely R image, run R, and make the files in your
current directory available to you. So you can load, test, and develop against
your current code and data locally, and save your changes. It’ll even save your
R workspace if you want it to.
For python, you might like to run an interactive ipython session:
opensafely exec python ipython
And similarly for stata:
opensafely exec stata-mp
You can also run cohortextractor directly, which can help testing study definitions:
opensafely exec cohortextractor --help
You can actually run any command you want by passing it as an argument.
For example, you can open an interactive bash shell in any of our images with, if you want to look around inside the image:
opensafely exec $IMAGE bash
For more information, you can run opensafely exec --help, or see the
documentation
Further work
We are always looking for ways to make developing with OpenSAFELY tools easier.
opensafely exec is a key feature to help acheive this. Using it as a base,
some other improvements we may be able to explore in future:
- An option to confgure Rstudio use the opensafely R image for OpenSAFELY projects
- Possibly running the graphical version of Stata
- VSCode integration to run your code automatically with
opensafely exec - Tooling to make testing your analysis code much easier.
New build of the OpenSAFELY R image
Updated: 15 February 2023
New version of the R image published
We have just published a new build of the R image. No action needed, but more information below!
What has not changed?
This is not a semantic change - the same versions of the same set of libraries are installed. However the way the image is built has been reworked, in order to address a number of shortcomings of the previous image, and provide a stonger base to build on in future.
What has changed?
We built the new image from a more up-to-date base. This means a number of core system libraries have been updated to newer versions. If you want to, you can see the list of system libraries that were updated. These are not R libraries, but rather some of the base C libraries used at a low level. Keeping them up to date is important for security and bug fixes.
In addition, we’ve moved from R 4.0.2 to 4.0.5 to pick up a slew of minor fixes.
What does this mean for me?
Hopefully, nothing! The next time you run opensafely pull, downloading the
R image may take a bit longer then normal. But otherwise, all your studies
should run as normal, locally and via jobs.opensafely.org.
We have tested the new image with variety of study code without issue, but if you do find something has changed, please contact tech support and let us know.
Who is to blame for this!?
Special thanks go to Tom Palmer from the University of Bristol for helping motivate and then exhaustively test the new build process.
- Pull request: https://github.com/opensafely-core/r-docker#123
Using compressed data files by default
Updated: 02 February 2023
Using compressed files is now the default recommendation in documentation and templates. On the backends, where datasets can be very large, using uncompressed files significantly slows execution and consumes more disk space.
The research-template has
been updated to generate csv.gz files from cohortextractor by default, and
the examples and Getting Started
documentation have been updated
to match.
In addition, recommendations for using compressed formats for further data files in python, R and Stata has been updated.
Because of the change in filename, if you have a workspace with a large amount of data in uncompressed CSV files, ask tech-support about moving to compressed CSVs, and we can help do this efficiently.
Improve handling of stale medication codelists in cohort-extractor
Updated: 06 January 2023
dm+d codes for Virtual Medicinal Products (VMPs) can change over time, with the result that codelists that contain VMP codes can become stale.
As a consequence, studies that use patients.with_these_medications()
with an old codelist may not have captured all the medication events that the author intended.
Until now, the mitigation has been to review and update codelists manually. With this update, cohort-extractor keeps stale medication codelists up-to-date by expanding the original codelist, so that it includes all current and previous codes of any VMPs in the codelist. This happens automatically.
Note that a codelist containing a VMP and all associated AMPs will still become stale with respect to new AMPs for that VMP are added to dm+d; this case is not currently handled automatically.
We will be contacting all study authors to explain the potential impact of using stale codelists more widely.
Add rounding instructions to disclosure documentations
Updated: 07 September 2022
Added instructions regarding rounding to our disclosure control documentation, including an example.
Added support for parameterised study definitions
Updated: 07 September 2022
Study definitions can now be “parameterised” so they accept values passed in by the action. This allows multiple cohorts with different properties to be generated by a single study definition.
Added support for querying by first and last day of the school year
Updated: 06 September 2022
Added support for querying by first and last day of the (English) school year.
Added resources management argument to opensafely run command
Updated: 31 August 2022
Users of the OpenSAFELY platform use the opensafely tool to run their code
locally prior to running against the real data.
This change adds some options to control the concurrency and memory usage of running actions, which should help users manage their local resource usage.
Added with_at_least_one_day_in_critical_care filter to admitted_to_hospital
Updated: 10 August 2022
Added ability to filter a patient’s hospital admissions to only those with at least one day in critical care.
Platform status page
Updated: 02 August 2022
We have rolled out a status page for the platform. Services on this page will be automatically updated if incidents occur.
Clarification about referrals data
Updated: 29 June 2022
We have clarified in the documentation that data on referrals is incomplete. Any codelists which include referral codes used with patients.with_these_clinical_events are unlikely to return complete data as it is largely held separately in a difficult-to-analyse format. We hope to offer an alternative data source for referrals in due course.
Oxford/AstraZeneca vaccination name change
Updated: 17 June 2022
The name of the Oxford-AstraZeneca vaccine has changed to COVID-19 Vaccine Vaxzevria 0.5ml inj multidose vials (AstraZeneca). To continue to access data for these vaccines, all users will need to update study definitions appropriately.
UK Renal Registry added to OpenSAFELY
Updated: 21 April 2022
Added support for querying the UK Renal Registry
ONS CIS added to OpenSAFELY
Updated: 05 April 2022
Added support for querying COVID-19 Infection Survey data provided by the Office for National Statistics.
Additional options for admitted_to_hospital
Updated: 08 March 2022
Added support for returning total_bed_days_in_period and total_critical_care_days_in_period from admitted_to_hospital.
ISARIC added to OpenSAFELY
Updated: 07 February 2022
Added support for querying the ISARIC (International Severe Acute Respiratory and Emerging Infection Consortium) dataset.
COVID-19 Therapeutics added to OpenSAFELY
Updated: 02 February 2022
Added support for querying the COVID-19 Therapeutics dataset.
Added ability to query a patient’s place_of_death
Updated: 28 January 2022
Added ability to query a patient’s place_of_death (TPP backend only).
Extended mean_recorded_value, added min_recorded_value and max_recorded_value
Updated: 27 January 2022
- Extended mean_recorded_value to support querying for mean recorded value across a full date range, as well as the most recent day of measurement.
- Added min_recorded_value to support querying for minimum recorded value.
- Added max_recorded_value to support querying for maximum recorded value.
- Pull request: opensafely-core/cohort-extractor#726
Added support for querying by first and last day of the NHS financial (recording) year
Updated: 24 January 2022
Added support for querying by first and last day of the NHS financial (recording) year.
Show how to extract cohortextractor output in compressed gzip (csv.gz) format
Updated: 06 September 2021
Show how to extract cohortextractor output in compressed gzip (csv.gz) format - recommended to be used in all studies to minimise file storage size. (However, CSVs can still be used where Stata is used for analysis).
This is now shown in all parts of documentation with cohortextractor steps.
Added support for users to provide their own dummy data
Updated: 21 July 2021
Added support for users to provide their own dummy data.
Documentation explaining how permissions work
Updated: 07 July 2021
Added a page in the docs explaining (briefly) how permissions work.
Ability to query a patient’s health care worker status in vaccination records
Updated: 11 June 2021
Add ability to query a patient’s health care worker status in vaccination records (TPP backend only)
OpenSAFELY CLI can now start a JupyterLab server
Updated: 18 January 2021
The OpenSAFELY command-line tool can now start a JupyterLab server. This allows you to easily develop code within the OpenSAFELY Python environment.
Added support for case-control studies
Updated: 14 January 2021
Added support for case-control studies
Subscribe to Platform News updates
Receive an email alert when the OpenSAFELY platform is updated.




